
What is Nuclear Magnetic Resonance - NMR
A very powerful analytical technique allowing chemists to identify even the most complex of structures.
Developed by chemists and physicists together it works by the interaction of magnetic properties of certain nuclei and their chemical environment.
This technique only works with atoms with an odd number of nucleons (protons and neutrons).
At A2 this will be applied to 11H and 136 C NMR.
Nuclear spin
All nucleons spin, and pair up just as electrons do.
|
|
|
|
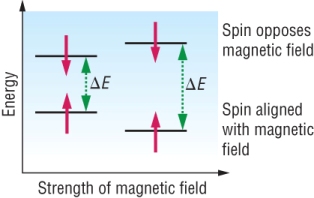 |
Resonance:
|
|
 |
Nuclear shielding and chemical shift:
1) Applied magnetic field
2) The weak magnetic fields generated from electrons surrounding the nuclei and nearby atoms (the environment)
|
|
|
Chemical shift, d :
|
|
|
Solvents for NMR spectroscopy:
|
|
|
Qu 1 - 2 P85
99% of any sample of carbon - 12C
1% of any sample of carbon - 13C
This 1% has an uneven number of nucleons, this means it will have a magnetic spin and be detected using NMR
Typical carbon - 13 chemical shifts:
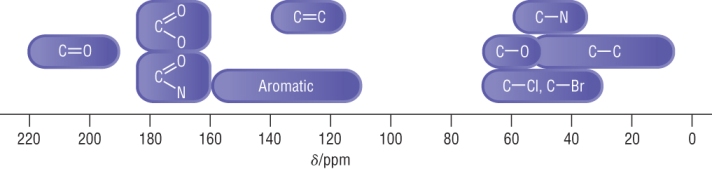
The chemical shift indicates the environments the 'carbons' are in.
An electronegative element causes a significant shift as carbon - 13 is sensitive to nuclear shielding.
The scale ranges 0 - 230, this means that each carbon is likely to have its own separate signal.
Values will vary with different solvents.
Interpreting carbon - 13 NMR spectra
3 things obtained from a carbon - 13 NMR is:
1) The number of different carbons
2) The carbon environment
3) The relative ratio of each of the types of carbons
|
|
|
Propan - 1 - ol:
|
Propan - 2 - ol:
|
Analysis of carbon - 13 NMR spectra
Making predictions:
Example 1: A carbonyl compound, C3H6O has the following C - 13 NMR:
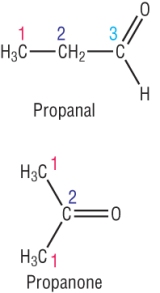 |
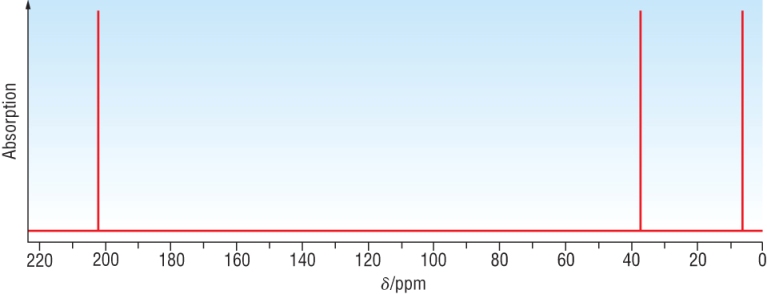 |
|
Example 2: An aromatic compound, C8H8O has the following C - 13 NMR:
|
|
||||||
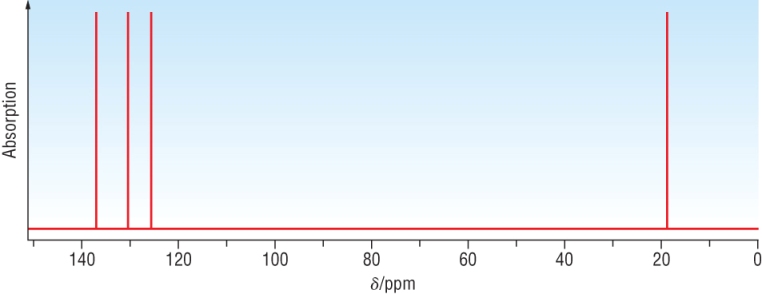 |
||||||
|
Possible structures: |
|
|||||
|
Carbon environments |
4 | 5 | 3 | 6 | ||
|
Aromatic environments |
3 | 4 | 2 | 4 | ||
Qu 1 - 2 P87 / Qu 1 P89
Proton NMR:
Is based around the 1H which is a single proton.
1H is much more abundant than 13C. 99.9% 1H to 1.1% 13C.
This means less needs to be used.
Proton NMR is done in the same way as 13C NMR and gives all the same information as 13C NMR but for protons.
In addition - it gives you information about adjacent protons (later)
Typical chemical shifts:
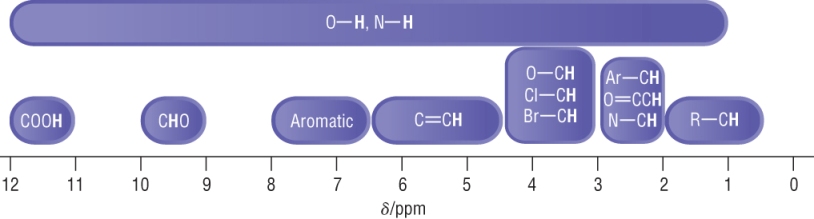
The scale is narrower which means some signals will overlap.
Actual chemical shifts can vary depending on environments.
The scale should be used as a rule of thumb.
Integration traces:
The area under the peak is proportional to the number of protons.
On the NMR spectrum, the spectrometer measures this and is recorded as an integration trace.
This is usually an integration line above the peak and can be measured for relative abundances.
Example: This is the proton NMR for C3H6O2
|
|
|
|
Qu 1 - 2 P91
Spin - spin coupling in proton NMR spectra
Spin - Spin coupling:
Splitting patterns are worked out by considering the effect that adjacent, chemically different hydrogen's have on another signal in a given environment.
The spin of the proton producing the signal is affected by each of the two forms of the adjacent hydrogen's (parallel and anti parallel).
One orientation enhances its field and the other reduces it.
We can work this out by calculating the various possible combinations of alignment of adjacent protons.
Theory:
The proton gives a signal by its magnetic field from its spin.
Its signal is influenced by adjacent protons (on neighbouring carbons).
Each proton will either spin in the same direction or the opposing direction.
This means that each adjacent proton either enhances the magnetic field or diminishes it.
There are 2 possibilities of equal chance per adjacent proton - enhancing or diminishing the magnetic field.
This splits the signal given by the proton
Analogy:
Imagine you had an opinion on something. If nobody influenced you, your opinion would be the same.
If another person had a view on the topic, they would either agree or disagree with you.
Their ideas would either enhance what you thought or diminish it.
There would be 2 possibilities of equal chance per person agreeing or disagreeing with you:
|
1 adjacent proton |
2 adjacent proton |
3 adjacent proton |
|
|
|
|
|
|
||
|
|
|
|
2 fields of equal intensity |
3 fields with an intensity of 1:2:1 | 4 fields with an intensity of 1:3:3:1 |
There is always an extra field than the number of adjacent protons - known as the n+1 rule:
n+1 rule:
|
n + 1 rule: Number of peaks = Number of different H's on adjacent atoms + 1 |
||||
| 1 Neighbouring H | 2 Peaks | DOUBLET | 1:1 | 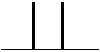 |
| 2 Neighbouring H | 3 Peaks | TRIPLET | 1:2:1 | 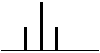 |
| 3 Neighbouring H | 4 Peaks | QUARTET | 1:3:3:1 | 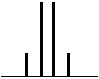 |
| 4 Neighbouring H | 5 Peaks | QUINTET | 1:4:6:4:1 | 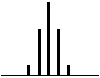 |
|
Signals for H in an O - H bond are unaffected by hydrogen's on adjacent atoms = singlet only |
||||
NOTE: Pascal's triangles
Just a note of interest. The signal peaks show the patterns described by Pascal's triangles:
The proton NMR spectrum of methyl propanoate:
There are 3 areas of protons - this will give 3 areas of signal:
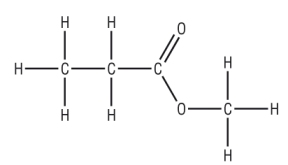
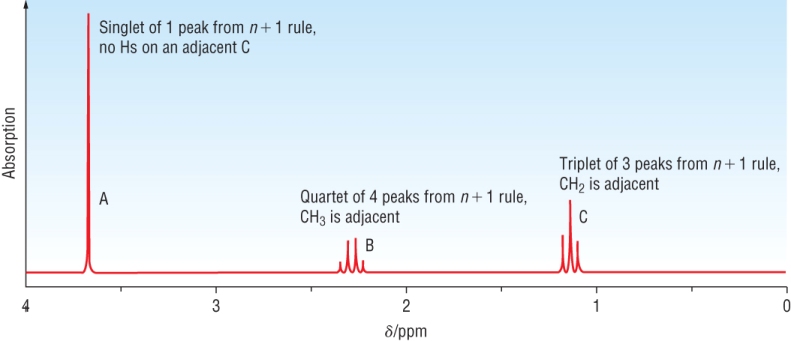
|
|
|
|
|
|
|
Qu 1 - 2 P93
NMR spectra of OH and NH protons
These are not only difficult to identify but can also confuse the rest of the spectra.
The reason for this is:
1) Peaks can appear over a wide range of chemical shifts
2) Signals are often broad
3) There is no splitting pattern (due to ease of proton exchange in OH / NH - not needed)
These signals can be removed by using heavy water, deuterium oxide, D2O
It is the same as water but the hydrogen's are replaced with deuterium.
Deuterium does not give a signal in NMR
Use of D2O
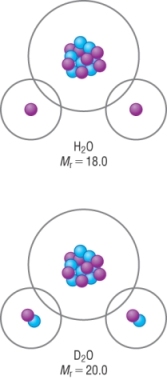 |
How D2O
is used: 1) An NMR is run as normal 2) A small amount of D2O is added to the mixture, shaken and a second NMR is run
How it works:
CH3CH2OH + D2O D CH3CH2OD + HOD
|
Example: NMR spectra of ethanol, (a) CH3CH2OH in water and (b) in D2O, CH3CH2OD
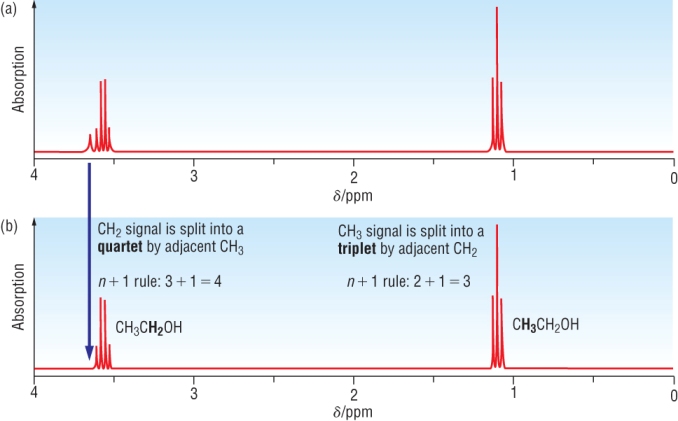
THE OH SIGNAL HAS DISAPPEARED
Splitting from -OH and -NH protons:
-OH and -NH peaks DO NOT split and DO NOT contribute to splitting
Hydrogen bonding between water (solvent) and -OH / -NH protons broaden the peak
Qu 1-2 P95
1) Using splitting patterns:
H - NMR of 2 isomers of C3H5ClO2: 1) CH3CHClCOOH and 2) ClCH2CH2COOH both run in D2O
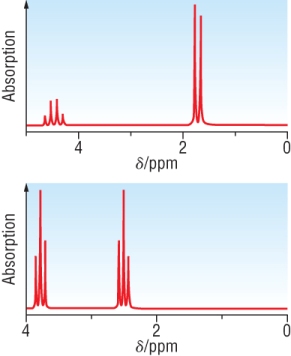 |
As it is run in D2O,
we do not need to worry about the COOH
signal.
|
2) Using splitting, integration and chemical shift:
H - NMR of 4 isomers of the ester, C4H8O2:
A) CH3CH2COOCH3 B) CH3COOCH2CH3 C) HCOOCH2CH2CH3 D) HCOOCH(CH3)2
2 of these esters are shown below, match the ester to the spectra:
.jpg) |
Chemical shift: | 1.1 | 2.1 | 3.6 |
|
|
.jpg) |
.jpg) |
||
|
Integration |
3 | 2 | 3 | |
| Splitting pattern | Triplet - signal adjacent to 2H's | Quartet - signal adjacent to 3H's | Singlet - signal adjacent to 0H's | |
| Interpretation | 3H's adjacent to 2H's | 2H's adjacent to 3H's | 3H's adjacent to 0H's | |
| Assignment | CH3CH2 | O=CCH2CH3 | O-CH3 | |
|
Put the assignments together: CH3CH2COOCH3
|
||||
49.jpg) |
Chemical shift: | 1.1 | 2.1 | 4.1 |
|
|
.jpg) |
.jpg) |
||
| Integration | 3 | 3 | 2 | |
| Splitting pattern | Triplet - signal adjacent to 2H's | Singlet - signal adjacent to 0H's | Quartet - signal adjacent to 3H's | |
| Interpretation | 3H's adjacent to 2H's | 3H's adjacent to 0H's | 2H's adjacent to 3H's | |
|
Assignment |
CH3CH2 | O=CCH3 | O-CH2CH3 | |
| Put the assignments together: CH3COOCH2CH3 | ||||
3) Protons adjacent on both sides:
The spectra below is for CH3CHClCH3
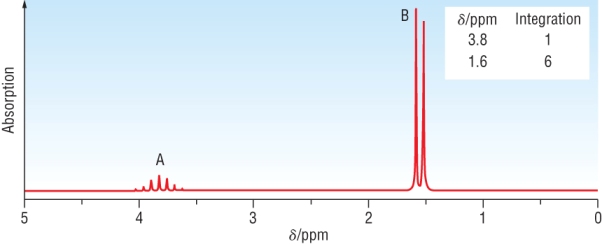 |
Chemical shift: | 1.6 | 3.8 |
|
|
.jpg) |
||
| Integration | 6 | 1 | |
| Splitting pattern | Doublet - signal adjacent to 1H's | Heptet - signal adjacent to 6 equivalent H's | |
| Interpretation | 6 equivalent H's adjacent to 1H's | 1H's adjacent to 6 equivalent H's | |
|
Assignment |
CH3CHClCH3 | CH3CHClCH3 | |
| Put the assignments together: CH3CHClCH3 | |||
4) Equivalent protons not split:
The spectra below is for ClCH2CH2Cl
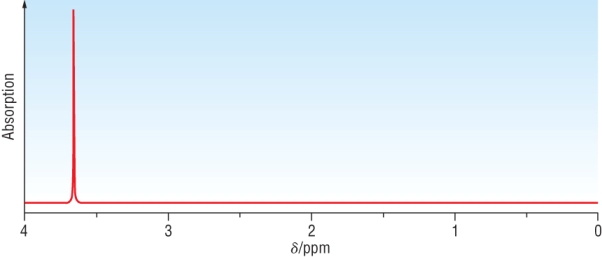 |
Chemical shift: | 3.8 |
.jpg) |
||
| Integration | 4 | |
| Splitting pattern | Singlet - signal adjacent to 2H's | |
| Interpretation | 2H's adjacent to 6H's | |
|
Assignment |
ClCH2CH2Cl x 2 | |
| Put the assignments together: ClCH2CH2Cl | ||
Qu 1 P97
Qu 1 - 4 P 99
| Mass Spectroscopy: |
|
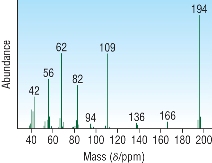 |
|
| IR spectroscopy: |
|
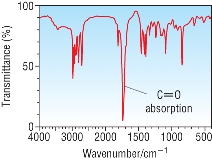 |
|
| NMR spectroscopy: | Carbon - 13 NMR:
Proton NMR:
|
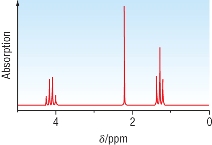 |
Worked example:
Chemical analysis has identified the empirical formula as C2H4O (Mr = 44
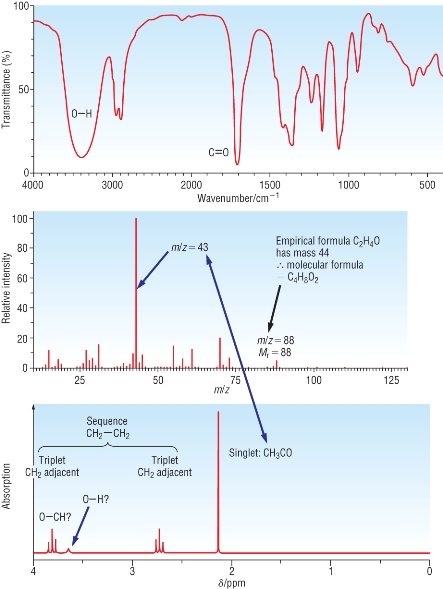 |
IR
spectra:
|
||||
Mass Spectra:
|
|||||
| NMR: | |||||
| Chemical shift: | 2.1 | 2.7 | 3.8 | 3.6 | |
.jpg) |
.jpg) |
.jpg) |
O-H can be in any region between 1.0 - 5.5 | ||
| Integration | 3 | 2 | 2 | 1 | |
| Splitting pattern | Singlet - signal adjacent to 0H's | Triplet - signal adjacent to 2H's | Triplet - signal adjacent to 2H's | Singlet | |
| Interpretation | 3H's adjacent to 0H's | 2H's adjacent to 2H's | 2H's adjacent to 2H's | O-H? | |
|
Assignment |
O=CCH3 | O=CCH2CH2- | O-CH2CH2 | -O-H | |
|
Put the assignments together:
|
|||||
Qu 1-2 P103 Qu 3-8 P105 Qu 2 - 8 P108/109